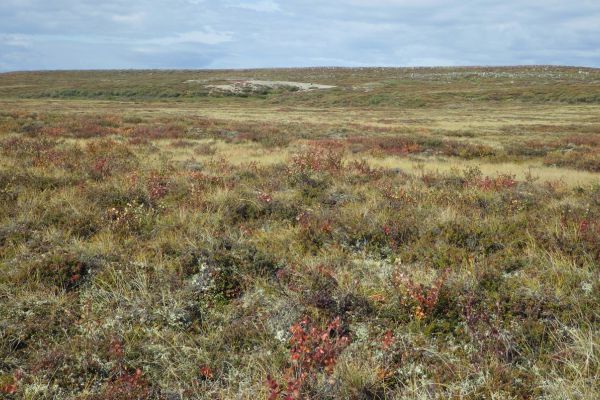
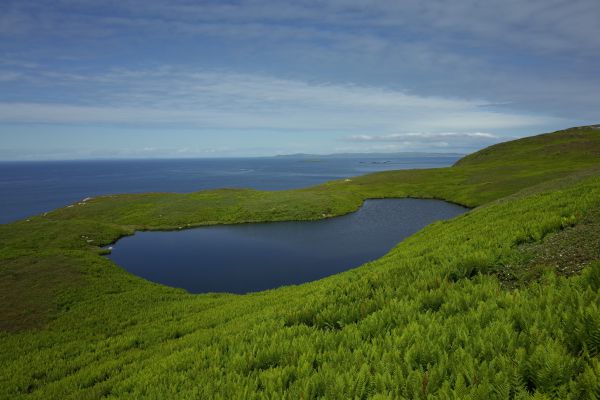
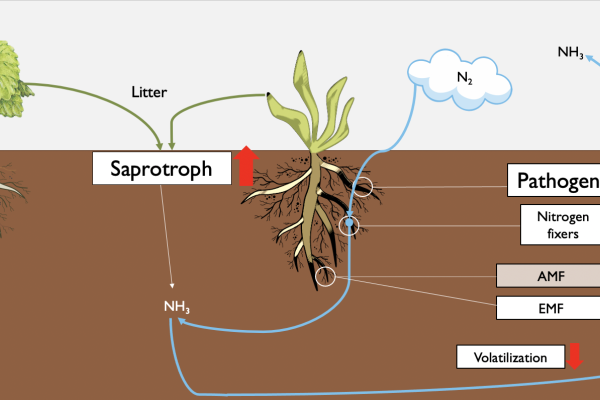
Explaining the “Greening” of the Arctic With Climate Change: Fungi May Help Birch Become More Dominant
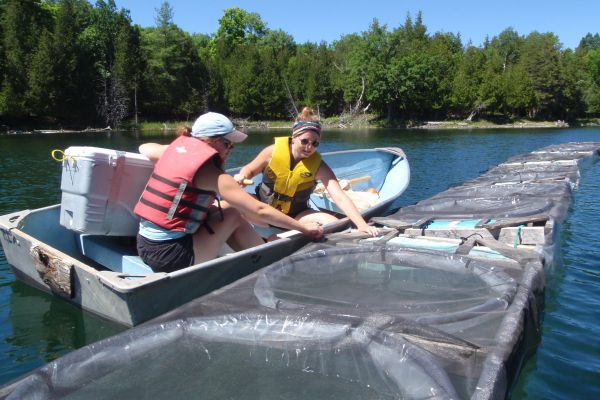
To model the impacts of climate change associated warming in Arctic soils, Queen’s University Biology researchers set up greenhouse and fertilizer amendment plots in the low Arctic tundra, 300 km NW of Yellowknife in the North West…
Current Water Quality Guidelines Across North America and Europe Do Not Protect Lakes From Salinization
Salinity is increasing in freshwater lakes worldwide, driven by the overuse of road salt, unsustainable agricultural practices, mining operations, and climate change. Increasing salinity is detrimental to biodiversity and overall lake…
Similar Zooplankton Responses to Low pH and Calcium May Impair Long-Term Recovery From Acidification
Uncontrolled, pervasive industrial emissions led to the acidification and destruction of biodiversity in North American and European lakes. Zooplankton, the pillars of freshwater ecosystems, are especially vulnerable to the associated pH…
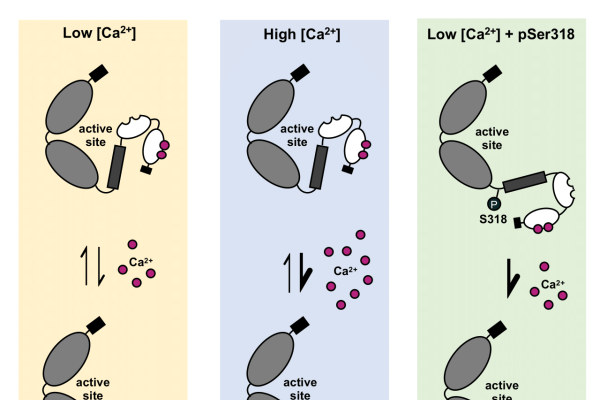
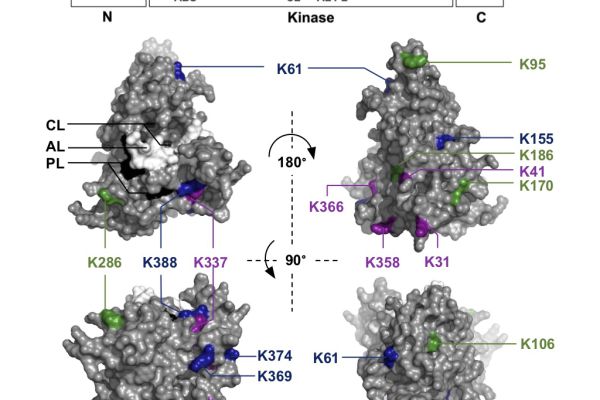
Phosphorylation-Dependent Subfunctionalization of the Calcium-Dependent Protein Kinase CPK28
Plants under stress may divert resources from growth and development to defence, leading to reduced yields for crops. These shifts in allocation are controlled by poorly known and complex signalling pathways involving phosphorylation and…
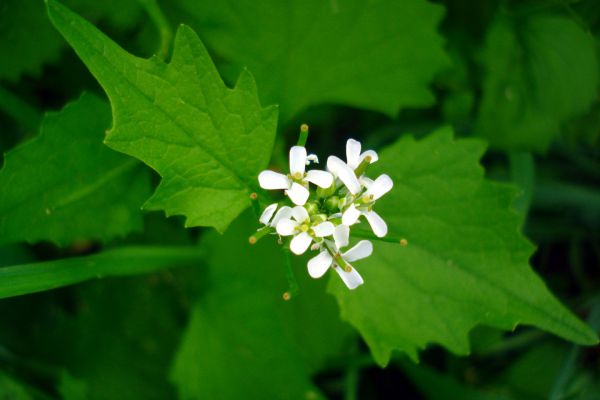
Genome Report: A Draft Genome of Alliaria Petiolata (Garlic Mustard) as a Model System for Invasion Genetics
Like many of Ontario’s invasive plants, garlic mustard ( Alliaria petiolata) is native to much of Eurasia and was probably introduced for medicinal and culinary purposes. Introduced to North America in the 19 th century, the species has…
The Moral Dilemma of the 21st Century
Advances in agriculture and medicine have given us now an overpopulated planet with severely impoverished ecosystem services — and a profound moral dilemma: how can we continue to respond to the still-growing need to feed more mouths and to…
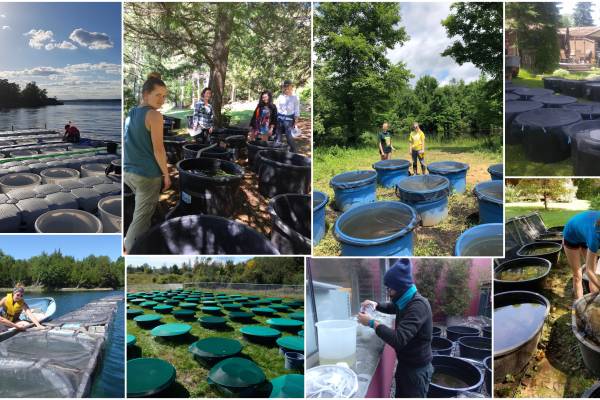
Effects of Chloride and Nutrients on Freshwater Plankton Communities
Over seven million tonnes of de-icing salt are dispersed on Canadian roads each winter leading to an increase in lake chloride concentrations that can be toxic to freshwater aquatic life. However, the current Canadian Water Quality…
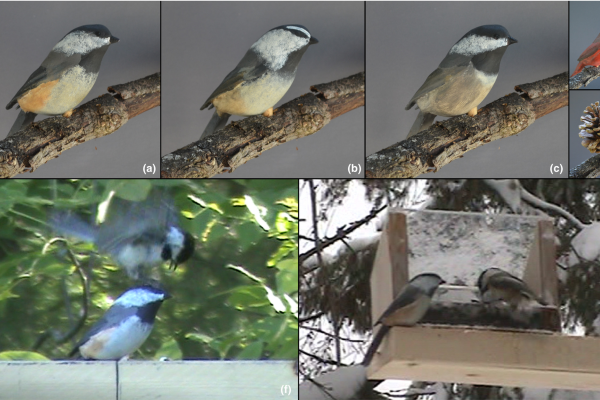
Experimental Tests of Selection Against Heterospecific Aggression as a Driver of Avian Colour Pattern Divergence
A healthy ecosystem relies on biodiversity which in turn relies on effective signal divergence to keep closely related, coexisting species distinct. For example, species-specific avian colour patterns allow birds to distinguish potential…
The Brachypodium Distachyon Cold-Acclimated Plasma Membrane Proteome Is Primed for Stress Resistance
Freezing damage can be particularly devastating with a single frost event capable of destroying billions of dollars worth of crops. To protect themselves, some plants enhance their freezing tolerance through an intricate process called cold…
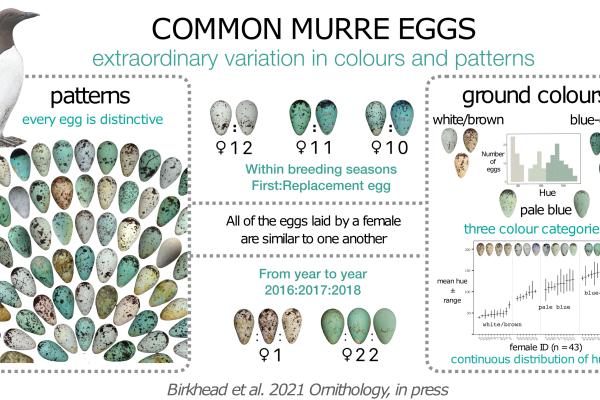
Exceptional Variation in the Appearance of Common Murre Eggs Reveals Their Potential as Identity Signals
The Common Murre, an avian artisan, crafts wondrously diverse eggs maculated with intricate designs that have enthralled naturalists and collectors for more than two centuries. Every egg is distinct; however, very little is known about the…
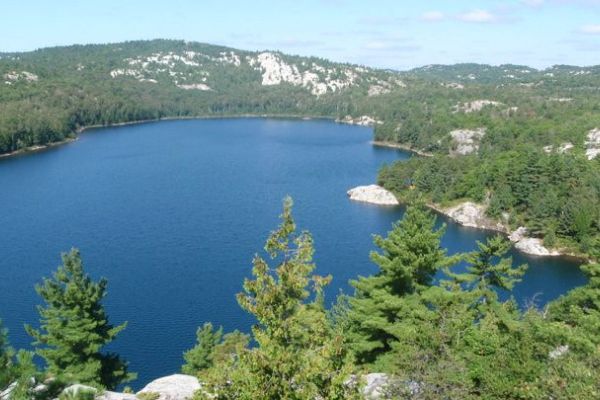
Investigating Holocene Climate Change in Boreal Northeast Ontario
Examining the past is an important tool for predicting the future. This is especially true when trying to predict the effects of global climate change on lakes and other freshwater ecosystems. Recently, researchers detailed changes in lakes…
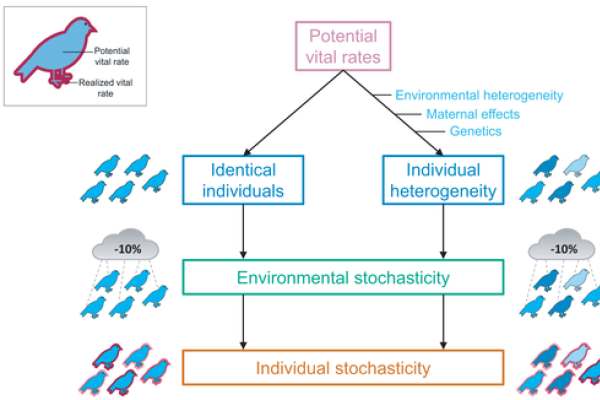
Demystifying Individual Heterogeneity
Individual vital rates, such as mortality and birth rates, are important determinants of life-histories and population trends. Models used to analyse population dynamics typically assume that individuals belonging to the same age or stage…
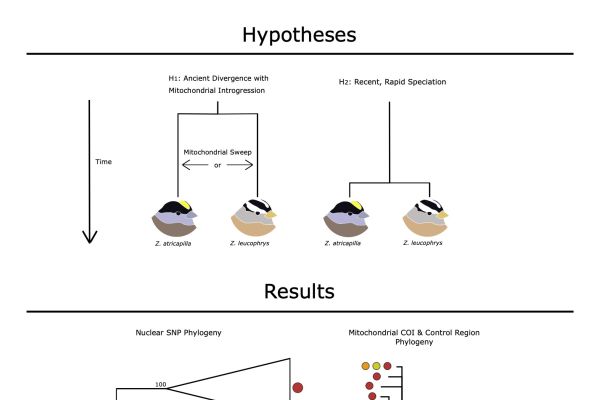
Cytonuclear Discordance in the Crowned-Sparrows, Golden-Crowned (Zonotrichia Atricapilla) and White-Crowned (Zonotrichia Leucophrys)
Golden-crowned and white-crowned sparrows have been hypothesized to have undergone rapid, and relatively recent (~50,000 years ago) speciation because they have nearly identical mitochondrial genomes. Nonetheless, the two species display…
Selection for Early Flowering Time in an Annual Plant (Yellow Rattle, Rhinanthus Minor) Occurs Regardless of an Elevational Gradient in Growing Season Length
Plant species that expand their geographic range typically adapt to new, local environmental factors via natural selection, becoming genetically and phenotypically differentiated from their source populations. Latitude and elevation are…
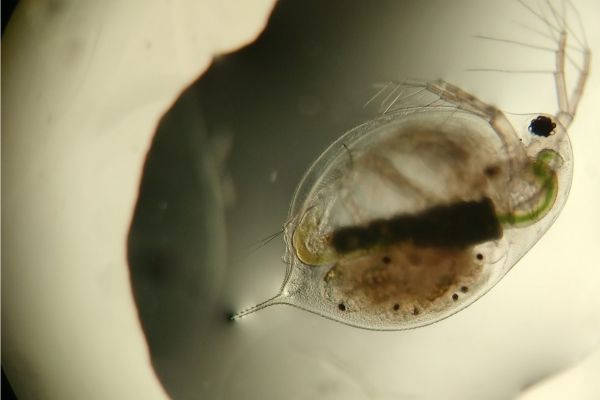
Drivers of Freshwater Zooplankton Communities
What factors determine the composition and abundance of freshwater communities? We know that smaller scale processes, such as soil or water chemistry, and broader scale processes, such as regional climate, are both important. However, few…
A Novel Allele of the Arabidopsis Thaliana MACPF Protein CAD1 Results in Deregulated Immune Signaling
Pathogen detection and response are critical components of defense against disease in all organisms. The Membrane Attack Complex/Perforin Family (MACPF) of proteins play an important role in immune responses in eukaryotic cells. There are…
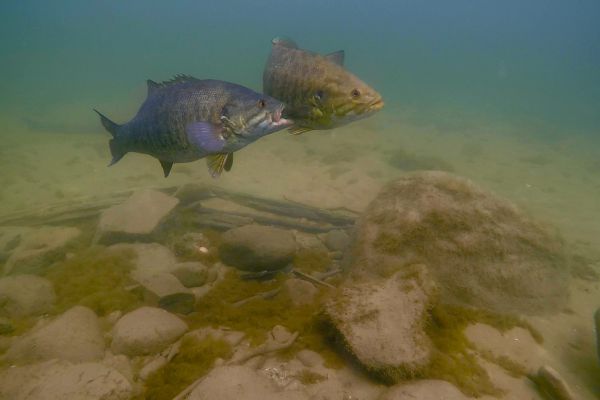
Angling Season Changes for Bass in Lake Ontario and the St. Lawrence River
Largemouth and smallmouth bass are very popular species for North American anglers, and angling for bass in the Great Lakes region provides economic benefits for many communities. In 2013, a change was implemented regarding the angling…
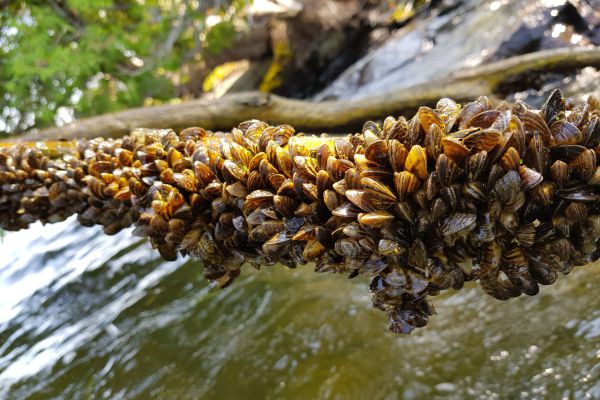
Are Current Watercraft Decontamination Measures Effective Against Aquatic Invasive Species?
Aquatic invasive species pose a major threat to all bodies of water, and the natural diversity that live in these ecosystems. Human activity is an important contributor to the spread of aquatic invasive species, often due to the improper…
The Effects of Weather on Avian Growth and Implications for Adaptation to Climate Change
Climate change is expected to cause plastic responses and evolutionary change across all taxa. For many bird species, climate change will alter weather patterns (such as temperature, rainfall, and wind), that will likely impact how young…
Species-Specific Conservation Strategies to Minimize Gray Ratsnake Road Mortality
Worldwide, biodiversity is declining as anthropogenic disturbance increases. A major threat to mammal, reptile, amphibian, and bird survival is road traffic. Many strategies to mitigate the effects of road traffic have been proposed…
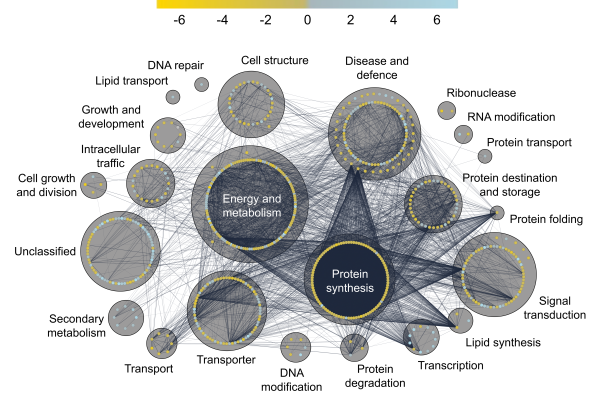
Identification of Ubiquitination Sites on Membrane-Associated Proteins in Arabidopsis
Protein phosphorylation and ubiquitination are two of the most commonly studied post-translational modifications of proteins in eukaryotes. While previous studies have recorded several ubiquitinated proteins in plants, few ubiquitinated…
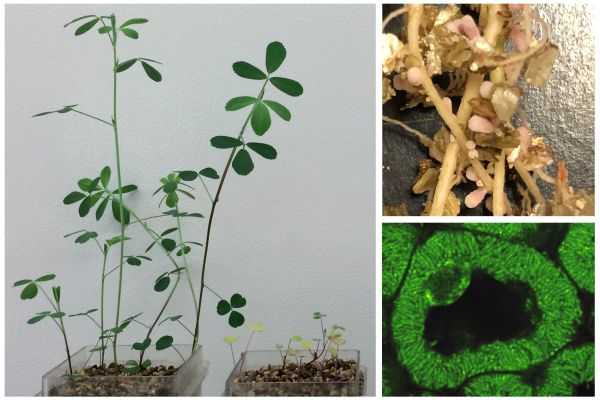
Investigating Transcriptional Responses in Plant-Rhizobia Symbioses
As the global demand for food production rises, a deep understanding of our crops – and their microbiomes – becomes essential. A plant’s microbiome includes a diverse suite of microbial species that are crucial in maintaining plant health…
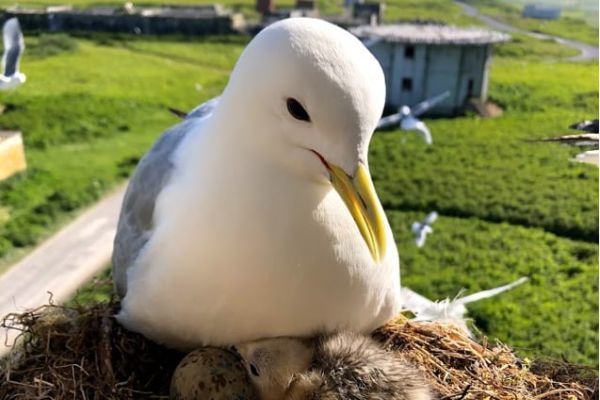
Seabird Population Decline Following European Settlement
Seabirds are important indicators of marine ecosystem health, and yet, we often lack long-term population data to inform conservation decisions. With ~70% of the world’s seabird populations in decline since the 1950s, long-term population…
Functional Shifts of Soil Microbial Communities Contribute to Invasion Success
Invasive plants can have devastating effects on native species and ecosystem processes. Garlic mustard ( Alliaria petiolata) is a problematic invader in North American deciduous forests. It produces chemicals that are thought to be…
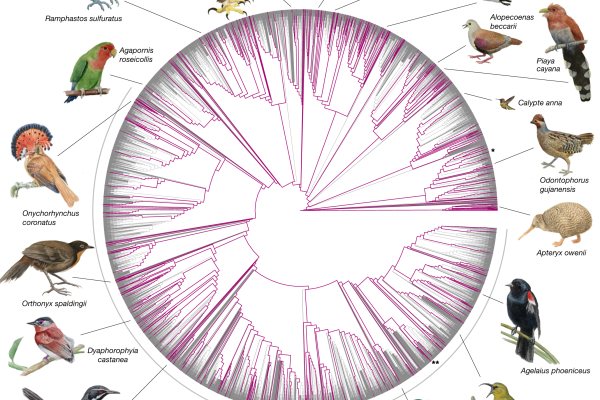
Dense Sampling of Bird Diversity Increases Power of Comparative Genomics
More than 10,000 species of birds are alive today, from the smallest hummingbird to the largest ostrich. Researchers have now captured a broad sampling of genomic information from across the avian tree of life. “Dense sampling of bird…
Celebrating Our Women Mentors
Queen’s Biology thanks our women mentors who are so instrumental to our development as scientists. Women mentors shape our scientific careers. They inspire us to pursue careers in biology, ensure we are confident and comfortable with…
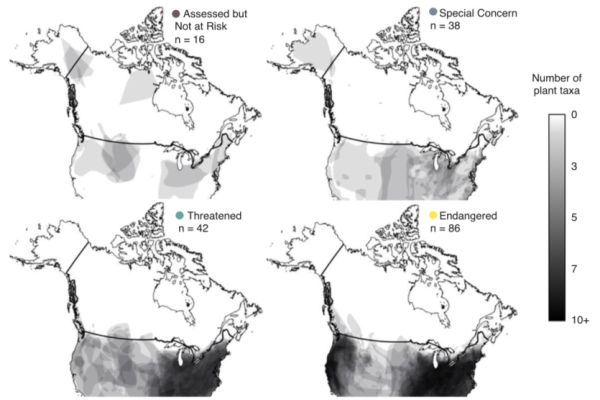
Range-Edge Plant Populations of High Conservation Priority Lack Adequate Habitat Protection and Research Effort
Southern Canada is home to many plants at the northern limits of their geographic ranges. These species are often rare and at-risk in Canada, though more broadly distributed south of the US border. While the conservation value of these…
What Causes Habitat Partitioning in Urban-Adapted Birds?
How important is plasticity versus evolutionary divergence for habitat partitioning in nature? In a recently published paper, Queen’s Biology Associate Professors Fran Bonier and Paul Martin, and former Queen’s Biology MSc student Kevin…